HBAT 2 (Hydrogen Bond Analysis Tool 2)#
A Python package to automate the analysis of potential hydrogen bonds and similar type of weak interactions in macromolecular structures from Protein Data Bank (PDB). HBAT supports both .pdb and .cif (mmCIF) file formats and uses a geometric approach to identify molecular interactions by analyzing distance and angular criteria.
Supported Interaction Types:
Hydrogen Bonds: Classical
N-H···O,O-H···O, and weakC-H···OinteractionsHalogen Bonds:
C-X···Ainteractions (X = Cl, Br, I)π Interactions:
X-H...πandC-X···πinteractions with aromatic rings (Phe,Tyr,Trp,His, etc.)π-π Stacking: Aromatic ring-ring interactions (parallel, T-shaped, offset)
Carbonyl Interactions:
n→π*interactions between carbonyl groupsn-π Interactions: Lone pair interactions with aromatic
πsystemsWater Bridges: Water-mediated hydrogen bond networks connecting protein/ligand residues
Ligand Interactions: Comprehensive detection of all interaction types between ligands and protein/nucleic acid residues
Announcement
HBAT 2 Web Interface is Live! Try it out at hbat-web.abhishek-tiwari.com

















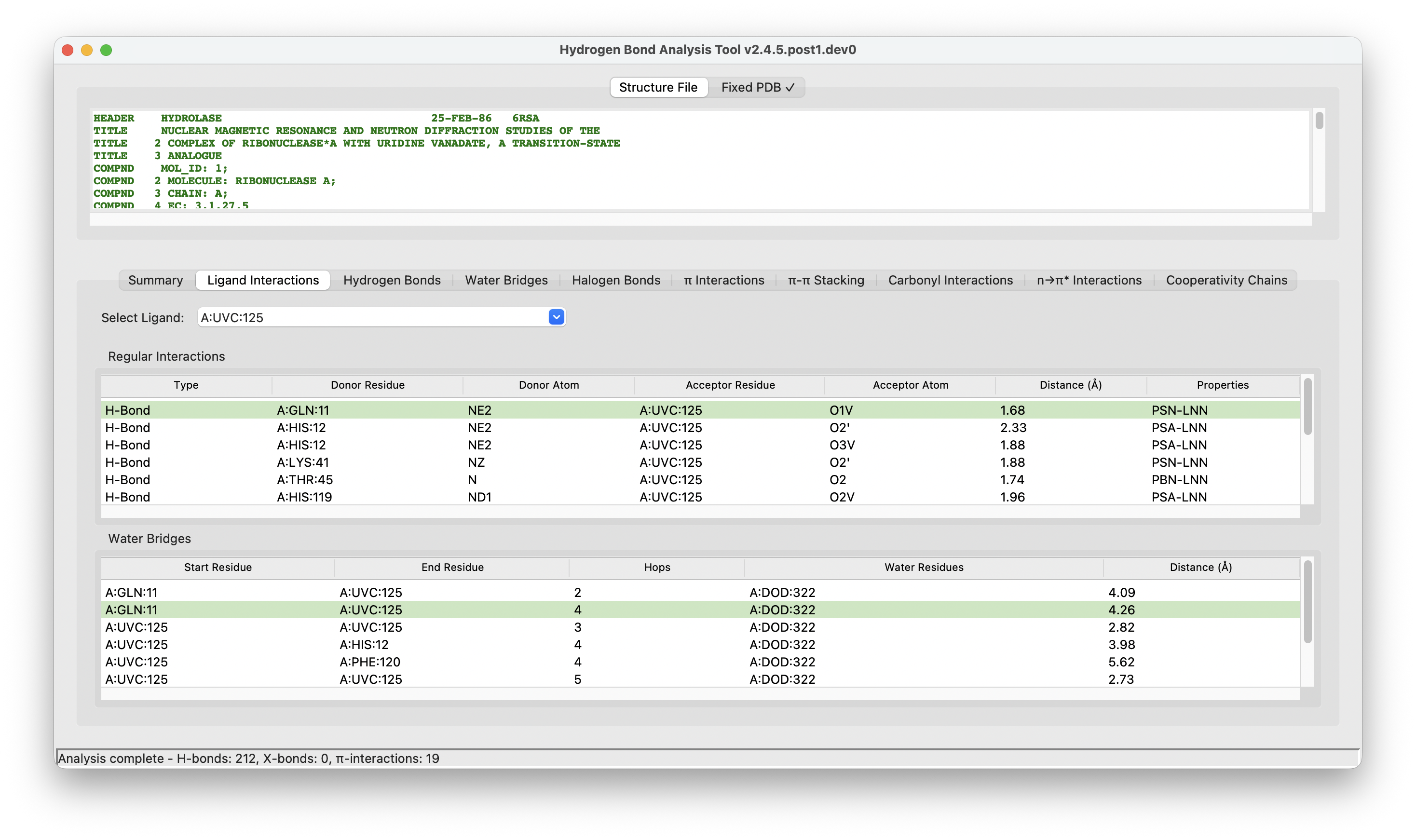
HBAT Desktop (Mac, Windows, Linux).#
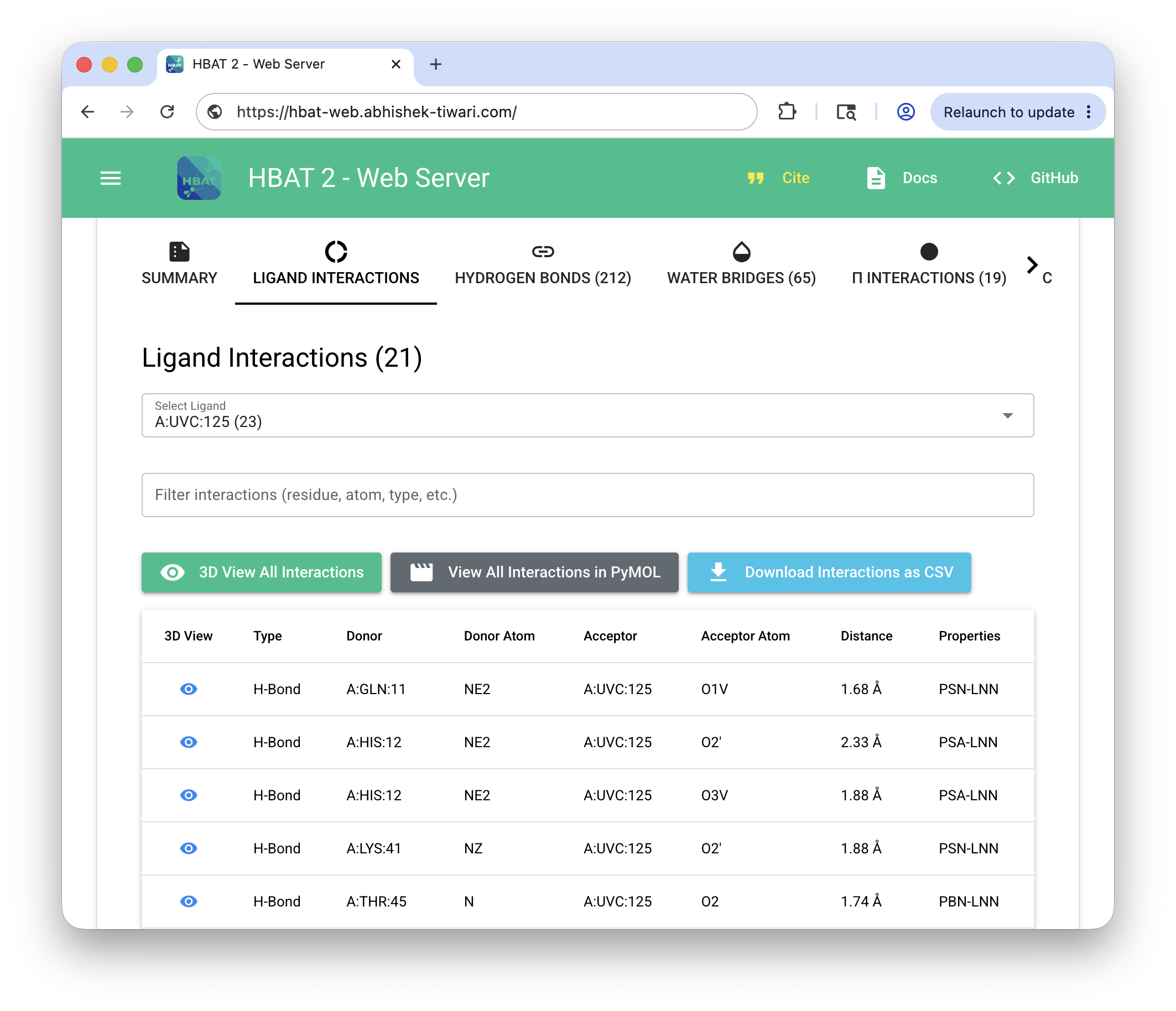
HBAT Web (https://hbat-web.abhishek-tiwari.com)#
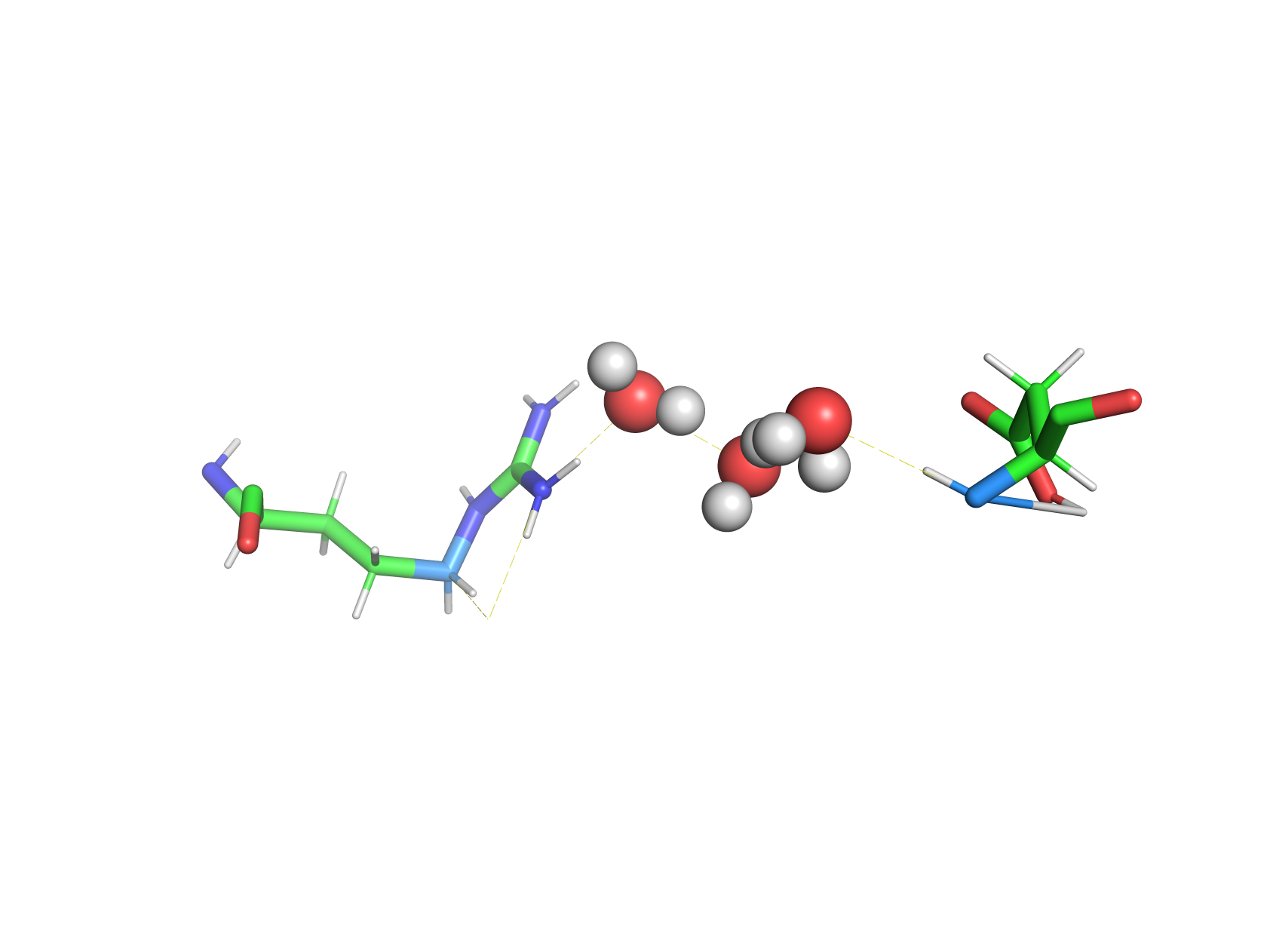
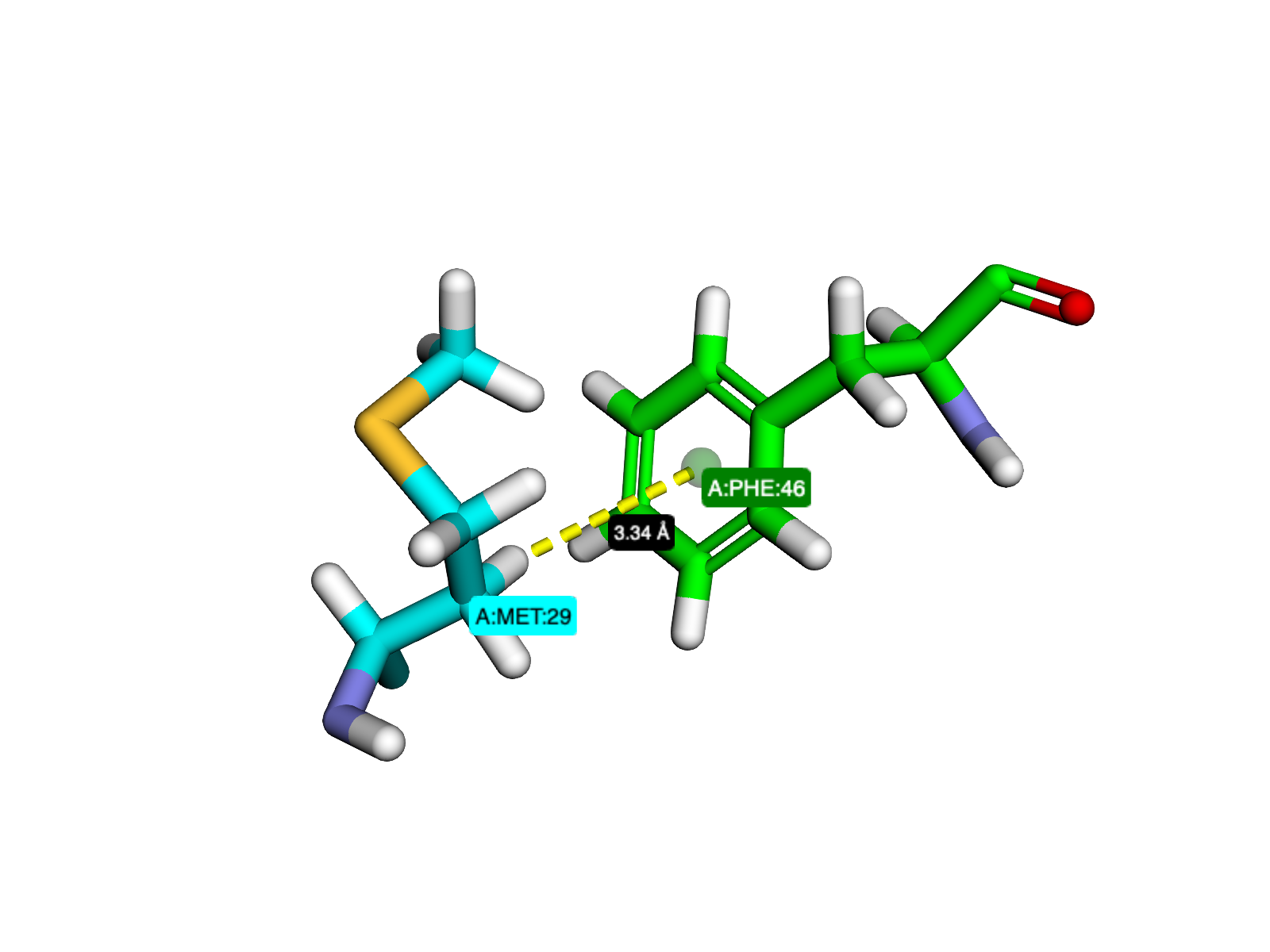
Visualizing interactions with HBAT Web using D3MOl and PyMOL (PDB Entry 6RSA)#
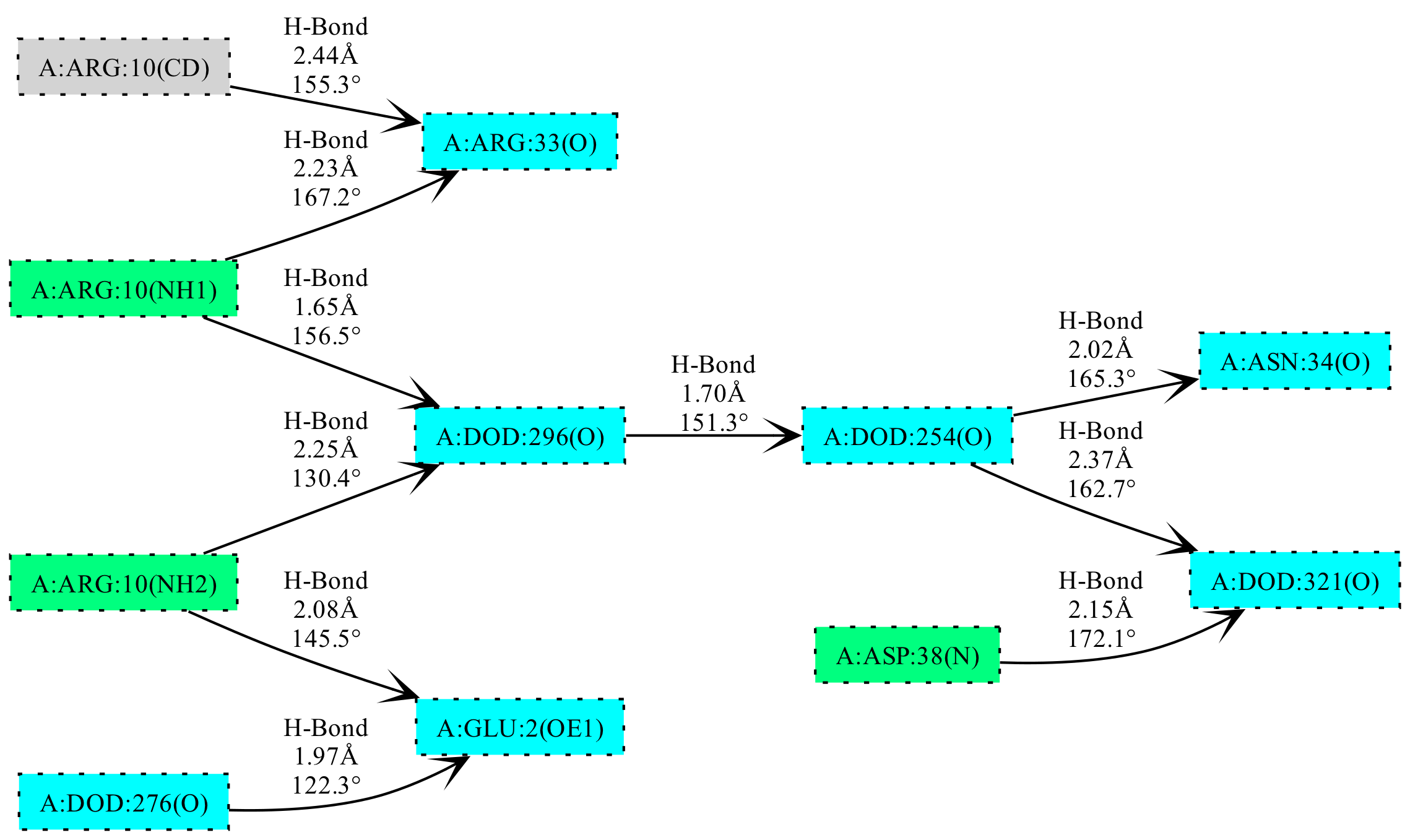
Cooperativity chain detection and visualization (PDB Entry 6RSA).#
Background#
HBAT 2 is a modern Python re-implementation of the original Perl-based tool developed by Abhishek Tiwari and Sunil Kumar Panigrahi. HBAT v1 can still be downloaded from SourceForge however Perl version is not maintained anymore.
Highlights of HBAT 2#
Detect and analyze potential hydrogen bonds, halogen bonds, π interactions, π-π stacking, carbonyl interactions, n-π interactions, water bridges, and ligand interactions
Automated PDB fixing with OpenBabel and PDBFixer integration
Support graphical (tkinter), command-line, and programming API interfaces
Use graphical interfaces for interactive analysis, CLI/API for batch processing and automation
Ligand interaction analysis with residue-specific visualization and filtering
Water bridge detection and analysis with bridge path visualization
Hydrogen bond network (potential cooperativity/anticooperativity chains and water-mediated hydrogen bond networks) visualization using NetworkX/matplotlib and GraphViz
Export hydrogen bond network visualizations to PNG, SVG, PDF formats
3D visualization of interactions using 3Dmol.js in Jupyter notebooks and HBAT web interface
Export and visualize interactions in PyMOL from HBAT web interface
Built-in presets for different structure types (high-resolution, NMR, membrane proteins, etc.)
Customizable distance cutoffs, angle thresholds, and analysis modes.
Multiple Output Formats: Text, CSV, and JSON export options
Optimized algorithms for efficient analysis of large structures
Cross-Platform: Works on Windows, macOS, and Linux.
Cite HBAT 2#
@article{tiwari_2026_hbat_arxiv,
author = {Tiwari, Abhishek},
title = {HBAT 2: A Python Package to analyse Hydrogen Bonds and Other Non-covalent Interactions in Macromolecular Structures},
year = 2026,
publisher = {arXiv},
doi = {10.48550/arXiv.2602.17712},
url = {https://arxiv.org/abs/2602.17712},
}
Tiwari, A. (2026). HBAT 2: A Python Package to analyse Hydrogen Bonds and Other Non-covalent Interactions in Macromolecular Structures. arXiv. https://doi.org/10.48550/arXiv.2602.17712
@article{tiwari_2026_hbat_chemrxiv,
author = {Abhishek Tiwari },
title = {HBAT 2: A Python Package to Analyse Hydrogen Bonds and Other Non-covalent Interactions in Macromolecular Structures},
publisher = {ChemRxiv},
year = {2026},
doi = {10.26434/chemrxiv.15000141/v1},
URL = {https://chemrxiv.org/doi/abs/10.26434/chemrxiv.15000141/v1},
eprint = {https://chemrxiv.org/doi/pdf/10.26434/chemrxiv.15000141/v1},
}
Tiwari, A. (2026). HBAT 2: A Python Package to analyse Hydrogen Bonds and Other Non-covalent Interactions in Macromolecular Structures. ChemRxiv. https://chemrxiv.org/doi/abs/10.26434/chemrxiv.15000141/v1
Cite HBAT 1.0 and 1.1#
@article{tiwari2007hbat,
author = {Tiwari, Abhishek and Panigrahi, Sunil Kumar},
doi = {10.3233/ISI-2007-00337},
journal = {In Silico Biology},
month = dec,
number = {6},
title = {{HBAT: A Complete Package for Analysing Strong and Weak Hydrogen Bonds in Macromolecular Crystal Structures}},
volume = {7},
year = {2007}
}
Tiwari, A., & Panigrahi, S. K. (2007). HBAT: A Complete Package for Analysing Strong and Weak Hydrogen Bonds in Macromolecular Crystal Structures. In Silico Biology, 7(6). https://doi.org/10.3233/ISI-2007-00337
User Guide